Discovery of Novel Adenosine Receptor Antagonists through a Combined Structure- and Ligand-Based Approach Followed by Molecular Dynamics Investigation of Ligand Binding Mode | Journal of Chemical Information and Modeling

Binding mode and protein-ligand interactions of 6-((4-fluorophenyl)... | Download Scientific Diagram

Exploration of Type II Binding Mode: A Privileged Approach for Kinase Inhibitor Focused Drug Discovery? | ACS Chemical Biology
Comparing Binding Modes of Analogous Fragments Using NMR in Fragment-Based Drug Design: Application to PRDX5 | PLOS ONE
Computer-guided binding mode identification and affinity improvement of an LRR protein binder without structure determination | PLOS Computational Biology
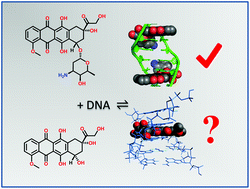
Investigations into the DNA-binding mode of doxorubicinone - Organic & Biomolecular Chemistry (RSC Publishing)

The crystal structure of the second Z-DNA binding domain of human DAI (ZBP1) in complex with Z-DNA reveals an unusual binding mode to Z-DNA | PNAS

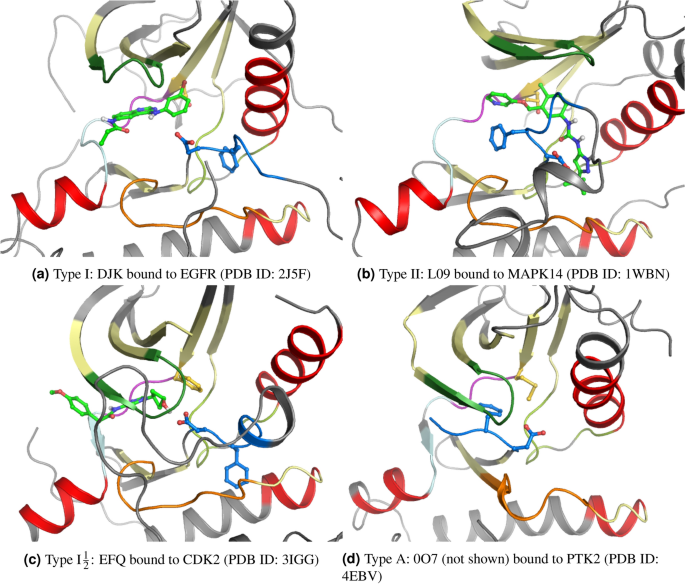
Exploration of Type II Binding Mode: A Privileged Approach for Kinase Inhibitor Focused Drug Discovery? | ACS Chemical Biology

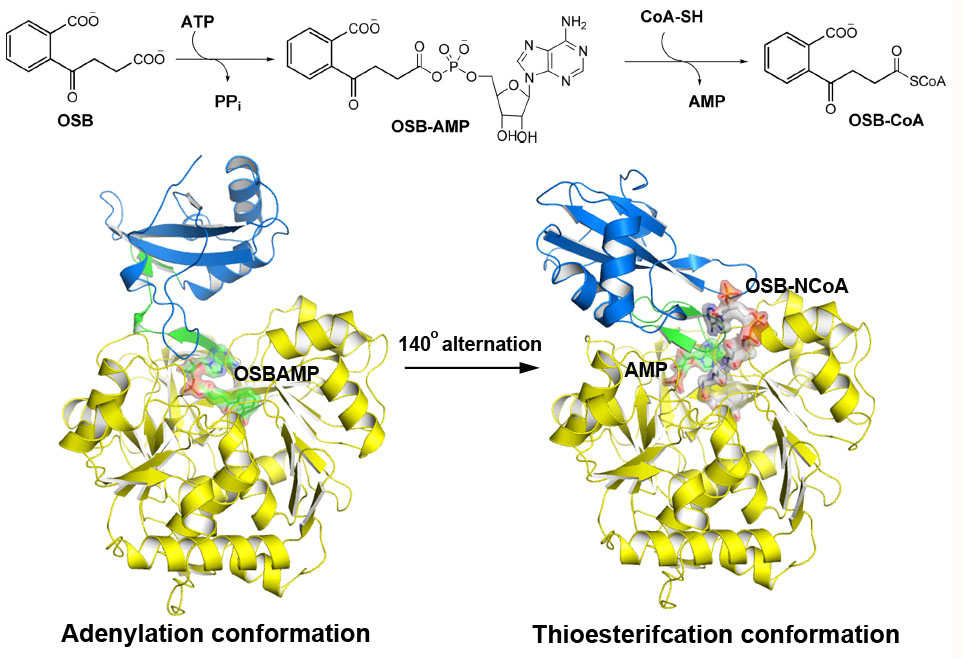
Universität Düsseldorf: Multiple Substrate Binding Mode-Guided Engineering of a Thermophilic PET Hydrolase
Prediction of kinase inhibitors binding modes with machine learning and reduced descriptor sets | Scientific Reports
The binding mode analysis of SI-4650. The binding modes of SI-4650 in... | Download Scientific Diagram
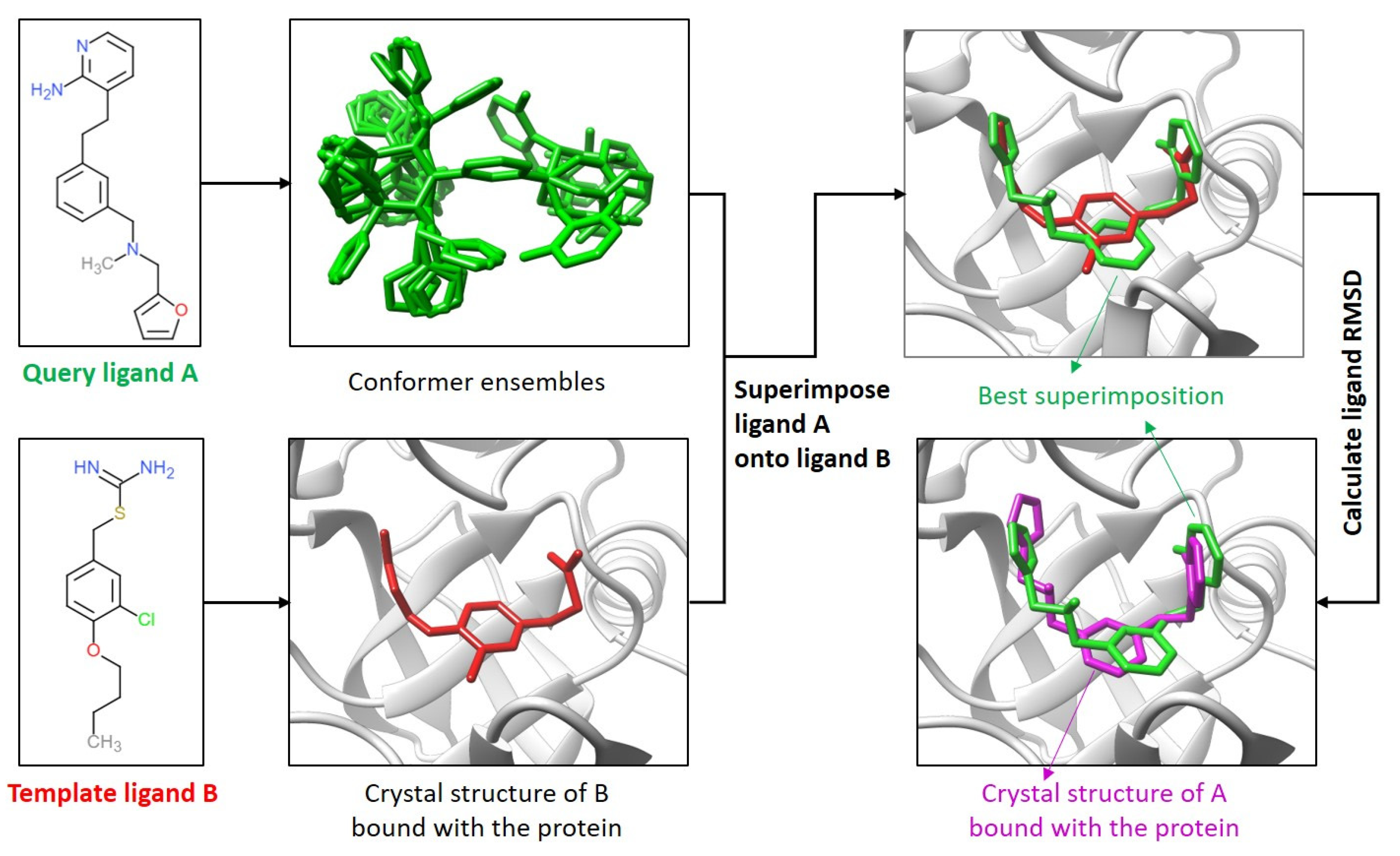
IJMS | Free Full-Text | Dissimilar Ligands Bind in a Similar Fashion: A Guide to Ligand Binding-Mode Prediction with Application to CELPP Studies

Prediction of ligand binding mode among multiple cross-docking poses by molecular dynamics simulations | SpringerLink

Proposed binding mode of 70551 into the N-Myc-Max DNA-binding domain... | Download Scientific Diagram

Binding mode of (A) lorlatinib and (B) Hit compound in the wild type... | Download Scientific Diagram

Binding mode comparison. (A) Binding mode of compound 10 in PB1(5). (B)... | Download Scientific Diagram
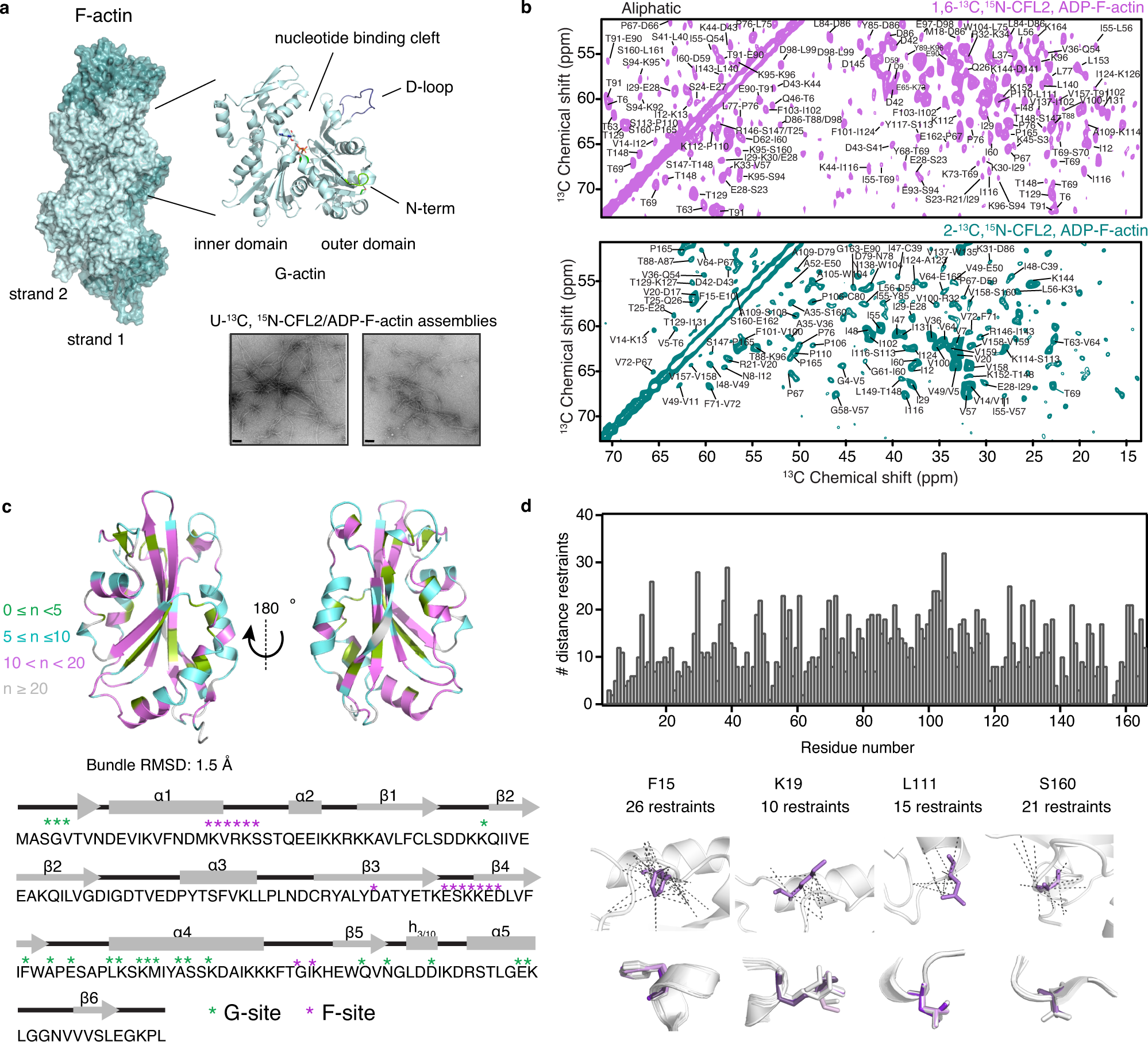
Magic angle spinning NMR structure of human cofilin-2 assembled on actin filaments reveals isoform-specific conformation and binding mode | Nature Communications

Binding modes of covalent small-molecule kinase inhibitors (CSKIs). (a)... | Download Scientific Diagram
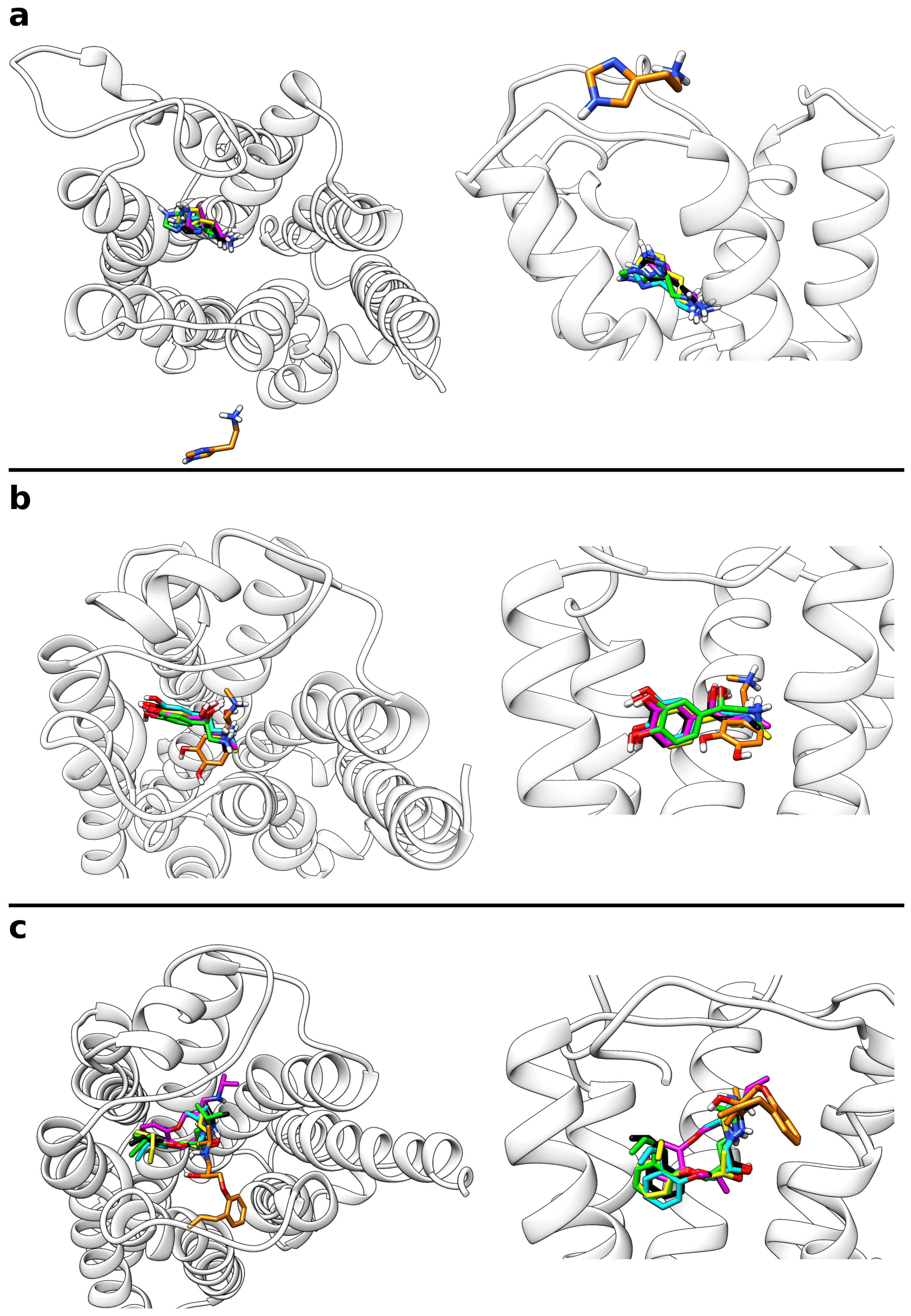
IJMS | Free Full-Text | A Metadynamics-Based Protocol for the Determination of GPCR-Ligand Binding Modes